-Search query
-Search result
Showing all 48 items for (author: estrozi & lf)

EMDB-19822: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+bromosterol (DOPC, DOPE, DOPS, bromo-ergosterol, PI(4,5)P2 35:20:20:15:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ
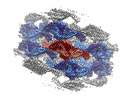

EMDB-18307: 
Native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Loewith RJ, Defosses A

EMDB-18308: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture -PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol 30:20:20:30)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18309: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/-sterol (DOPC, DOPE, DOPS, PI(4,5)P2 50:20:20:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18310: 
Helical reconstruction of yeast eisosome protein Pil1 bound to membrane composed of lipid mixture +PIP2/+sterol (DOPC, DOPE, DOPS, cholesterol, PI(4,5)P2 35:20:20:15:10)
Method: helical / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18311: 
Compact state - Native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18312: 
Stretched state - Native eisosome lattice bound to plasma membrane microdomain
Method: single particle / : Kefauver JM, Zou L, Desfosses A, Loewith RJ

EMDB-18043: 
Helical structure of the influenza A virus ribonucleoprotein-like
Method: helical / : Chenavier F, Estrozi LF, Zarkadas E, Ruigrok RWH, Schoehn G, Ballandras-Colas A, Crepin T
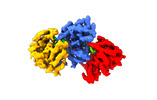
EMDB-18044: 
Focused reconstruction of influenza A RNP-like particle
Method: helical / : Chenavier F, Estrozi LF, Zarkadas E, Ruigrok RWH, Schoehn G, Ballandras-Colas A, Crepin T

EMDB-15627: 
Tomogram of the Mimivirus genomic fiber
Method: electron tomography / : Villalta A, Schmitt A, Estrozi L, Quemin ERK, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Gruenewald K, Abergel C

EMDB-15628: 
Tomogram of the Mimivirus genomic fiber
Method: electron tomography / : Villalta A, Schmitt A, Estrozi L, Quemin ERK, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Gruenewald K, Abergel C

EMDB-15629: 
Tomogram of the Mimivirus genomic fiber
Method: electron tomography / : Villalta A, Schmitt A, Estrozi L, Quemin ERK, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Gruenewald K, Abergel C

EMDB-15630: 
Tomogram of the Mimivirus genomic fiber
Method: electron tomography / : Villalta A, Schmitt A, Estrozi L, Quemin ERK, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Gruenewald K, Abergel C

EMDB-13641: 
Structure of the Mimivirus genomic fibre asymmetric unit
Method: single particle / : Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
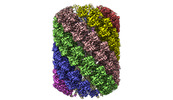
EMDB-14353: 
Structure of the Mimivirus genomic fibre in its compact 6-start helix form
Method: helical / : Villalta A, Schmitt A

EMDB-14354: 
Structure of the Mimivirus genomic fibre in its compact 5-start helix form
Method: helical / : Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C

EMDB-14355: 
Structure of the Mimivirus genomic fibre in its relaxed 5-start helix form
Method: helical / : Villalta A, Schmitt A

EMDB-11179: 
Head of Semi-jumbo phage RP13
Method: single particle / : Neumann E, Kawasaki T, Effantin G, Estrozi L, Chatchawankanphanich O, Yamada T, Schoehn G

EMDB-11180: 
Head reconstruction of full jumbo phage XacN1
Method: single particle / : Neumann E, Kawasaki T, Effantin G, Estrozi L, Chatchawankanphanich O, Yamada T, Schoehn G

EMDB-11275: 
MreC
Method: helical / : Estrozi LF, Contreras-Martel C

EMDB-11178: 
Jumbo Bacteriophage RSL2 - Full icosahedral capsid
Method: single particle / : Neumann E, Kawasaki T, Effantin G, Estrozi L, Chatchawankanphanich O, Yamada T, Schoehn G

EMDB-10926: 
Structure of jumbo coliphage phAPEC6 capsid
Method: single particle / : Wagemans J, Tsonos J, Holtappels D, Fortuna K, Hernalsteens JP, De Greve H, Estrozi LF, Bacia-Verloop M, Moriscot C, Noben JP, Schoehn G, Lavigne R

EMDB-10929: 
3D structure of bacteriophage phAPEC6 tail
Method: single particle / : Wagemans J, Tsonos J, Holtappels D, Fortuna K, Hernalsteens JP, De Greve H, Estrozi LF, Bacia-Verloop M, Moriscot C, Noben JP, Schoehn G, Lavigne R

EMDB-20086: 
In situ structure of rotavirus VP1 RNA-dependent RNA polymerase (TLP)
Method: single particle / : Jenni S, Salgado EN

EMDB-20087: 
In situ structure of rotavirus VP1 RNA-dependent RNA polymerase (DLP)
Method: single particle / : Jenni S, Salgado EN

EMDB-20088: 
In situ structure of rotavirus VP1 RNA-dependent RNA polymerase (TLP_RNA)
Method: single particle / : Jenni S, Salgado EN

EMDB-20089: 
In situ structure of rotavirus VP1 RNA-dependent RNA polymerase (DLP_RNA)
Method: single particle / : Jenni S, Salgado EN

EMDB-0326: 
Aeromonas hydrophila ExeD
Method: single particle / : Contreras-Martel C, Farias Estrozi L

EMDB-0327: 
Vibrio vulnificus EpsD
Method: single particle / : Contreras-Martel C, Farias Estrozi L

EMDB-4179: 
Combined solid-state NMR, solution-state NMR and EM data for structure determination of the tetrahedral aminopeptidase TET2 from P. horikoshii
Method: single particle / : Gauto DF, Estrozi LF, Schwieters CD

EMDB-4039: 
Maltose binding protein genetically fused to dodecameric glutamine synthetase
Method: single particle / : Coscia F, Petosa C
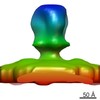
EMDB-4006: 
Structure of wt PFV glycoprotein by cryo-electron tomography
Method: subtomogram averaging / : Effantin G

EMDB-4007: 
Structure of iFuse PFV glycoprotein by cryo-electron tomography
Method: subtomogram averaging / : Effantin G

EMDB-4008: 
Structure of the wt PFV glycoprotein from the iNAB Gag mutant by cryo-electron tomography
Method: subtomogram averaging / : Effantin G

EMDB-4010: 
Structure of a hexagonal assembly of the wt PFV glycoprotein from the iNAB Gag mutant by cryo-electron microscopy
Method: single particle / : Effantin G

EMDB-4011: 
Structure of a hexagonal assembly of the wt PFV glycoprotein from the iNAB Gag mutant by cryo-electron microscopy
Method: single particle / : Effantin G

EMDB-4012: 
Structure of a hexagonal assembly of the PFV glycoprotein from the iFuse Env mutant by cryo-electron microscopy
Method: single particle / : Effantin G

EMDB-4013: 
Structure of a hexagonal assembly of the PFV glycoprotein from the iFuse Env mutant by cryo-electron microscopy
Method: single particle / : Effantin G

EMDB-2690: 
Inside rotavirus at 7 A resolution
Method: single particle / : Estrozi LF

EMDB-2691: 
Inside rotavirus at 7 A resolution
Method: single particle / : Estrozi LF

EMDB-2823: 
Structure determination of feline calicivirus virus-like particles in the context of a pseudo-octahedral arrangement
Method: single particle / : Burmeister WP, Buisson M, Estrozi LF, Schoehn G, Billet O, Hannas Z, Sigoillot-Claude C, Poulet H

EMDB-2628: 
Structural similarity of secretins from Type II and Type III Secretion Systems
Method: single particle / : Tosi T, Estrozi LF, Job V, Guilvout I, Pugsley AP, Schoehn G, Dessen A

EMDB-2629: 
Structural similarity of secretins from Type II and Type III Secretion Systems
Method: single particle / : Tosi T, Estrozi LF, Job V, Guilvout I, Pugsley AP, Schoehn G, Dessen A

EMDB-2100: 
Location of the dsRNA-dependent polymerase, VP1, in rotavirus particles
Method: single particle / : Estrozi LF, Settembre EC, Goret G, McClain B, Zhang X, Chen JZ, Grigorieff N, Harrison SC

EMDB-1762: 
Cryo-EM structure of the E. coli translating ribosome in complex with SRP and its receptor
Method: single particle / : Estrozi LF, Boehringer D, Shan SO, Ban N, Schaffitzel C

EMDB-1752: 
3D reconstruction of the rotavirus VP6 trimer using the Fast Projection Matching (FPM) algorithm.
Method: single particle / : Estrozi LF, Navaza J

EMDB-1463: 
Three-dimensional structure of canine adenovirus serotype 2 capsid
Method: single particle / : Schoehn G, El Bakkouri M, Fabry CM, Billet O, Estrozi LF, Le L, Curiel DT, Kajava AV, Ruigrok RW, Kremer EJ

EMDB-1462: 
Three-dimensional structure of canine adenovirus serotype 2 capsid
Method: single particle / : Schoehn G, El Bakkouri M, Fabry CM, Billet O, Estrozi LF, Le L, Curiel DT, Kajava AV, Ruigrok RW, Kremer EJ
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
